Andrew Collette, Research Scientist with IMPACT, HDF Guest Blogger
“…HDF5 is that rare product which excels in two fields: archiving and sharing data according to strict standardized conventions, and also ad-hoc, highly flexible and iterative use for local data analysis. For more information on using Python together with HDF5…”

An enormous amount of effort has gone into the HDF ecosystem over the past decade. Because of a concerted effort between The HDF Group, standards bodies, and analysis software vendors, HDF5 is one of the best technologies on the planet for sharing numerical data. Not only is the format itself platform-independent, but nearly every analysis platform in common use can read HDF5. This investment continues with tools like HDF Product Designer and the REST-based H5Serv project, for sharing data using the HDF5 object model over the Internet.
What I’d like to talk about today is something very different: the way that I and many others in the Python world use HDF5, not for widely-shared data but for data that may never even leave the local disk… The HDF5 object model is extremely simple. There are only three categories of object:
- datasets that store numerical data;
- groups, hierarchically organized containers which store datasets and other groups; and
- attributes, named bits of metadata which can be attached to groups and datasets.
When a data product needs to be widely shared, the groups, datasets and attributes are strictly arranged according to a convention. Some of these conventions are very well-defined; for example, HDF-EOS is used for data products from NASA’s Earth Observing System and specifies a long list of required attributes and groups to represent various earth-science constructs.
Over the past several years, HDF5 has emerged as the mechanism of choice for storing large quantities of numerical data in Python. Two high-profile packages now exist for interfacing with HDF5: PyTables, a table-oriented product used by the pandas analysis package, among others; and h5py, a general-purpose interface. Writing data with either package is very straightforward, as both use the facilities of the Python language to handle the routine, boilerplate operations for you. For example, using the h5py package, here’s how to open an HDF5 file, store an array, and clean up by closing the file:
with h5py.File("myfile.hdf5") as f:
f["data"] = myarray
Because of the inherent flexibility of HDF5, the lack of a required schema, absence of a database and the associated configuration headaches, combined with the very low verbosity of Python code, the vast majority of files created with either h5py or PyTables do not follow any established convention. Neither are they totally unstructured. Rather, such files follow ad-hoc domain- or application-specific conventions, which are often made up on the fly.
Here’s an example. I work as a research scientist at the University of Colorado’s IMPACT facility. Funded by NASA, we operate a large machine which simulates micrometeorites and dust particles in space, by accelerating microscopic iron spheres to extreme speeds – up to 200,000 miles per hour. Data from the machine emerges in a mixture of formats; some in HDF5 and following a loosely-documented lab-specific convention, and some in old binary formats from our digital oscilloscopes.
Rarely does one go from raw machine data to final, publication-ready plots. In between, the data is processed and reduced… bad or corrupted data is thrown away, timestamps are synchronized, and the remaining information is searched and analyzed for scientific value.
For even a modest experimental run, the volume of data is far too large to fit in memory, and the required computation too expensive to repeat every time some detail of the analysis changes. An intermediate form of the data is required, and it’s here that the combination of Python and HDF5 really shines.
Here’s an example of one such file, from an experiment I performed a few weeks ago. It was created from the raw data by a 150-line Python script. Each iron grain impact is represented by a group containing a few datasets. Each group has a standard set of attributes I made up on the spot for use in my analysis:
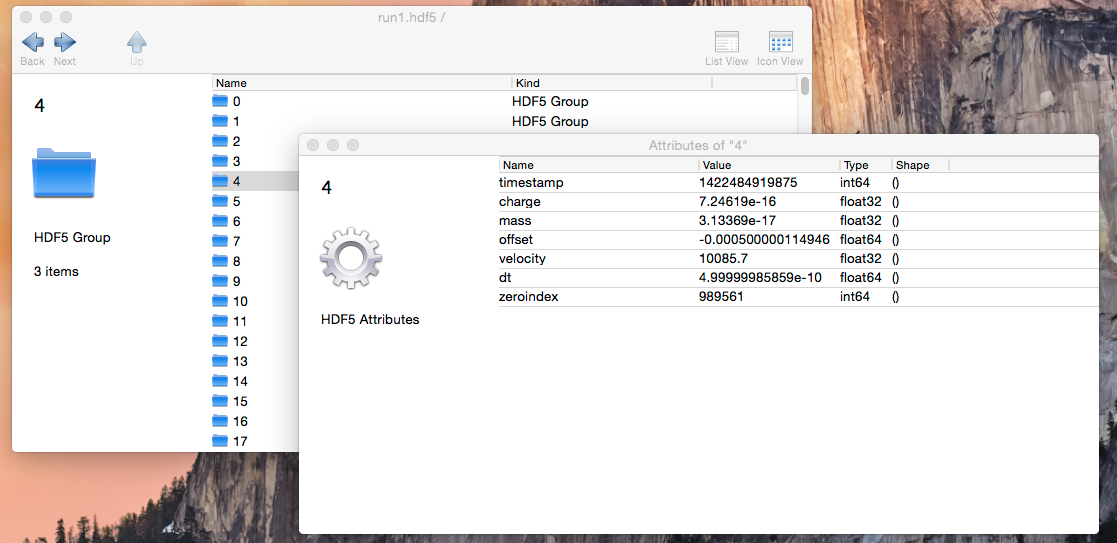
The only consumer for these files at the moment is my analysis code. But that’s OK… if I were to share this file, anyone using an HDF5-compatible analysis environment would be able to make sense of it. Because HDF5 is self-describing, anyone using Python, IDL, MatLab or any of a dozen other environments could simply open up the file and poke around. They would recognize that each group had, for example, a “mass” attribute (which represents the iron grain’s mass for this particular impact). Writing code to pick out, say, only masses above a certain threshold would be trivial.
So HDF5 is that rare product which excels in two fields: archiving and sharing data according to strict standardized conventions, and also ad-hoc, highly flexible and iterative use for local data analysis. For more information on using Python together with HDF5, here are a couple of links to get you started:
• The h5py project: http://www.h5py.org
• PyTables: http://www.pytables.org
• Pandas analysis environment (which uses HDF5): http://pandas.pydata.org
Editor’s Note: Thank you, Dr. Collette, for a great article! More information about the accelerator can be found here.